The politics of time with Carolyn Smith

As CEO of the Cambridge Crystallographic Centre, Senior Member and alumnus Jürgen Harter (PhD Chemistry, 1998) is leading the effort to find drugs to target COVID-19.
Jürgen Harter has plenty of reasons to look back fondly on his time as a PhD student at Wolfson (Chemistry, 1998). Not only was he active in various University societies, and enjoyed the social activities at College (especially the College bar) but it's also where he met his future wife, Dr Catherine Ngozi Harter (PhD Psychiatry, 1998). They married in the summer of 2003.
The couple have maintained their connections to Wolfson, with a circle of friends that include Wolfson alumni all over the world, and they both became Senior Members in 2003. With plenty of international travel required by his work, Jürgen hasn’t spent as much time at College as he would have liked, but in recent years the couple have been drawn back to attend lectures and join the other Senior Members for dinner. But that will have to wait. For now, Jürgen is active in the effort to combat COVID-19.

Once he completed his PhD, Jürgen took on a series of roles in the life sciences industry, including business development and digital transformation. In July 2018 he became CEO of the Cambridge Crystallographic Data Centre (CCDC). During the Coronavirus pandemic, the Centre has redirected some of its resources to help researchers worldwide who are looking for potential treatments for COVID-19.
The CCDC was established in 1965 and is a registered not-for-profit organisation with close links to the University of Cambridge. At its heart is the mammoth Cambridge Structural Database (CSD), a databank of nearly 1 million organic and metal organic crystal structures. The structures can be viewed as three-dimensional models and include information about their chemical and physical properties.They are of interest to academics and researchers in numerous industries, including pharmaceutical, agrochemical and biotechnology.
The Centre has also developed software tools to help researchers explore the CSD. Amongst them is the Discovery Suite, which is run by an in-house scientific team and includes a tool called GOLD, which stands for ‘Genetically Optimised Ligand Docking’. GOLD allows researchers to find binding sites on proteins where small molecules known as ligands can ‘dock’ or attach themselves. When a ligand successfully docks, it creates a bioactive signal that can be measured, identifying the binding site as a potential target for drugs.
Although GOLD had been developed for some time, it's focus had been on lead optimisation; making highly accurate predictions on a small number of promising drug candidates, says Jürgen. But with the increase in computing power offered by the cloud, the potential to scale up the application of GOLD to screen millions of compounds in high-throughput docking became apparent, and resources were invested to develop this capability. This turned out to be of benefit when COVID-19 appeared.
“I became aware of Coronavirus straight away, and I knew we had a research role to play with all of the chemistry data and software that we have,” says Jürgen. “I was in Singapore in December at the Asian Crystallographic Conference and it was pretty clear what was going on in Wuhan. Colleagues from other Asian countries were already at heightened alert; they knew what to do when an epidemic strikes. I came back to Cambridge and said, we have to get to work on this right away.”
In January, Chinese scientists determined the crystal structure of the main protease of SARS-CoV-2 called MPro, an enzyme that the virus needs to replicate itself. CCDC scientists have helped others in assessing over 4,000 potential compounds that have been suggested by worldwide experts. The results of these endeavours have been made freely available. The team have also been working on generating new drug ideas to submit to this effort in collaboration with researchers at The University of Cambridge and Sheffield University. In this context, millions of synthetically accessible potential compounds are generated in silico, docked and scored. The most promising candidates generated from this process are then further assessed and submitted for assay in laboratory tests. Many other groups worldwide have used GOLD for similar efforts.
With 20% of the team at the Centre devoted to the COVID-19 pandemic, there are several other projects underway. CCDC researchers are scouring the literature to track any small molecule drugs of interest, and making these available to researchers in academia and industry. And other researchers are looking at different proteins on the SARS-CoV-2 virus.
Jürgen says, “We are also looking at several other structural interpretations of the Mpro protein, which demonstrates its structural plasticity.” The structure of the virus was first described by a team of Chinese researchers: it is very similar to a related structure from the previous SARS outbreak. Using this structure, a team at the Diamond Light Source, the UK’s national synchrotron facility in Oxfordshire, undertook a fragment screening program to observe locations for the binding of small fragments which can in turn inform drug design. Others too have published structures of the virus bound to potential inhibitors. Due to the urgency of the pandemic, researchers have rightly pushed this information out quickly — but this has led to some inaccuracies. The Centre has been keen to work with its collaborators to provide improved models.
“If the structure of the virus is not correctly described then people will not be doing the right research and will get off on the wrong track,” he says.
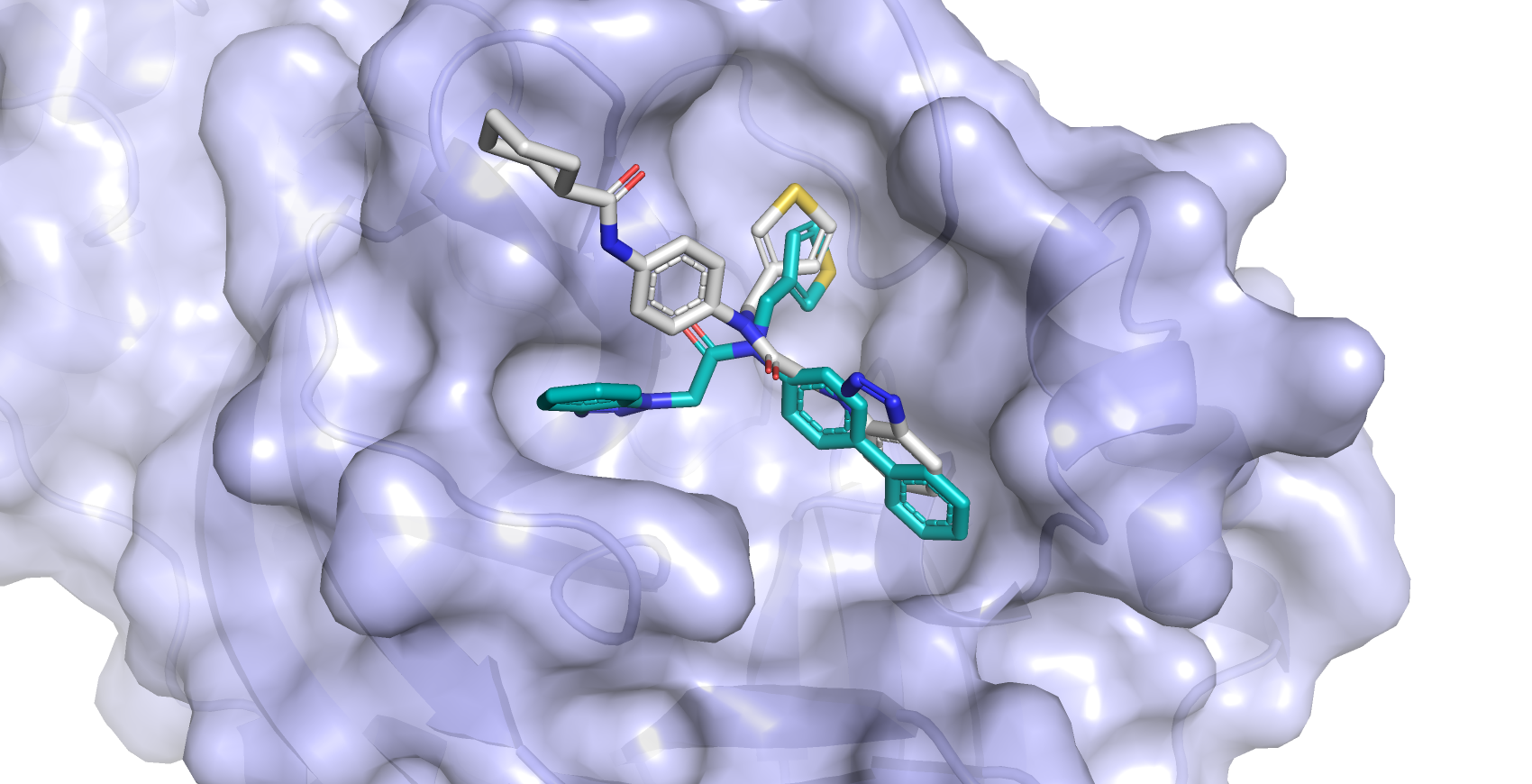
Artificial intelligence also contributes to the accuracy of the Centre’s data. “We’ve been using machine learning for about 10 years with our various software packages,” says Jürgen. “GOLD uses genetic algorithms to self-optimise the way the ligand-protein docking happens. The more it docks, the better the results over time.”
The Centre has joined the PostEra COVID Moonshot startup, an international group of scientists from academia and industry trying to develop drugs to combat COVID-19. PostEra also uses machine learning to accelerate the development and testing of compounds based on the molecules of interest. Moonshot is taking submissions of molecules from around the world and then all of those suggestions are run through GOLD to see if they are good candidates to enter the work flow for drug development.
“We figure this is an international effort to defeat the virus and we’ve got to use what we have to throw at the problem. They need our data to do a better job with the AI. The combination of AI and the good data will ultimately beat this thing,” says Jürgen. “We are trying to beat the brains of medicinal chemists. The creativity of the human mind is unmatched but the volume of what the AI algorithms can process is better. So I think the machines will be winning with this one.”
Since the pandemic began, Jürgen has taken his team of 75 out of their offices in Central Cambridge and they are all working from home, along with their colleagues in the US.
“It’s been quite a transition to home-working model,” says Jürgen. “But we haven’t really missed a beat because we can remote-control the centre and we are using the University High Performance Computer Cluster to run our software.”
In addition to his efforts with COVID, Jürgen and Catherine are also home schooling, although he admits that it is Catherine who is the primary teacher (“I kind of hover in and out of the dining room when my kids are sitting with their laptops.”)
The couple are looking forward to coming back to College when the regulations permit, and Jürgen is especially looking forward to talking chemistry with Jane Clarke.
“You have a great new President in Jane Clarke – well, not so new any more,” he says, “And I remember her from the Chemistry Department. What I like about Wolfson is that in its essence it hasn’t changed since I was a student. It still has what I always loved about it, being such an international community with such a great intellectual atmosphere.”
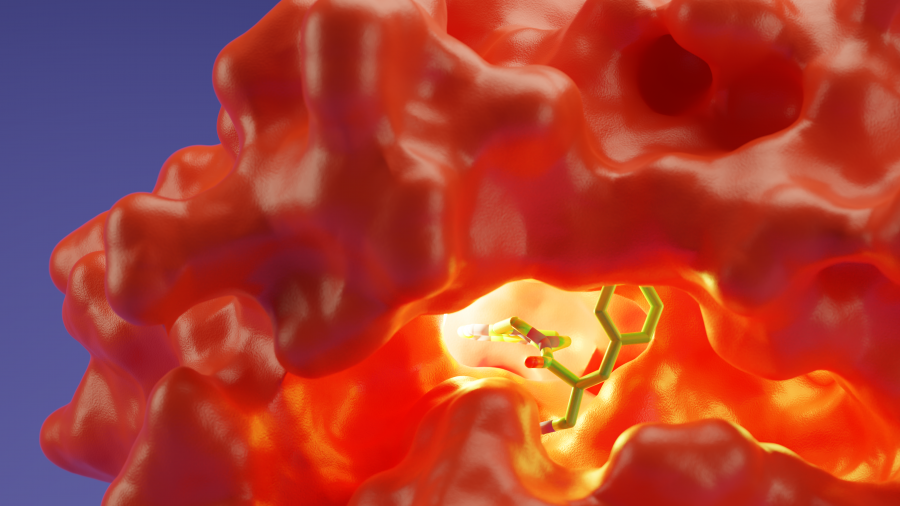
Main image: Key binding points for inhibitors against the COVID-19 M-Pro protease highlighted in yellow. As identified by CCDC sponsored PhD students Mihaela Smilova (Oxford) and Peter Curran (Cambridge and UCB)







